A role for tubulin in cellular quality control and proteostasis
Abstract
Microtubules (MTs), stiff rods built up from tubulin dimers, form a cytoskeletal network whose structure, behaviour, and function have been extensively investigated, mainly from a mechanical perspective. Here, we describe a role for tubulin in the cellular stress response. We overexpressed tubulin dimers in a controlled fashion in 293F cells. Despite the engagement of autoregulation, a mechanism that degrades tubulin-encoding mRNAs when tubulin levels are high, a surplus of tubulin and microtubules is detected in overexpressing cells. This leads to altered microtubule behaviour, mitotic problems, deregulation of the cell cycle, and replication stress. Surprisingly, we also observe proteostasis defects in tubulin overexpressing cells, which we attribute to mitochondrial stress-related translation attenuation. Conversely, tubulin and microtubules are downregulated as part of the response to oxygen or glutamine deprivation. Together, our data link tubulin levels, and hence autoregulation, to cellular quality control and proteostasis. We propose that competitive interactions with key partners, including the mitochondrial protein import and general translation machinery, underlie the tubulin-mediated control of cellular homeostasis.
A role for tubulin in cellular quality control and proteostasis
Sreya Basu¹²*, Nuo Yu¹, Riccardo Viscusi¹, Wouter Dof³, Mirjam van den Hout⁴, Wilfred F.J. van IJcken⁴, Karel Bezstarosti³, Dick H. W. Dekkers³, Jeroen Demmers³, Niels Galjart¹²*
¹Department of Cell Biology, Erasmus University Medical Center Rotterdam, The Netherlands ²Department of Clinical Pharmacy, Erasmus University Medical Center Rotterdam, The Netherlands ³Center for Proteomics, Erasmus University Medical Center Rotterdam, The Netherlands ⁴Center for Biomics, Erasmus University Medical Center Rotterdam, The Netherlands
*Corresponding authors: s.basu@erasmusmc.nl ; n.galjart@erasmusmc.nl
bioRxiv preprint doi: https://doi.org/10.64898/2026.04.06.716648
Abstract
Microtubules, stiff rods built up from tubulin dimers, form a cytoskeletal network whose structure, behaviour, and function have been extensively investigated, mainly from a mechanical perspective. Here, we describe a role for tubulin in the cellular stress response. We overexpressed tubulin dimers in a controlled fashion in 293F cells. Despite the engagement of autoregulation, a mechanism that degrades tubulin-encoding mRNAs when tubulin levels are high, a surplus of tubulin and microtubules is detected in overexpressing cells. This leads to altered microtubule behaviour, mitotic problems, deregulation of the cell cycle, and replication stress. Surprisingly, we also observe proteostasis defects in tubulin overexpressing cells, which we attribute to mitochondrial stress-related translation attenuation. Conversely, tubulin and microtubules are downregulated as part of the response to oxygen or glutamine deprivation. Together, our data link tubulin levels, and hence autoregulation, to cellular quality control and proteostasis. We propose that competitive interactions with key partners, including the mitochondrial protein import and general translation machinery, underlie the tubulin-mediated control of cellular homeostasis.
Introduction
Microtubules (MTs), one of the three major cytoskeletal networks of eukaryotic cells, are dynamic structures composed of heterodimers of α and β tubulin. These building blocks assemble in a head-to-tail arrangement into MTs, with one end of the MT (the minus end) exposing alpha-tubulin, and the other end (the plus-end) beta-tubulin. Soluble tubulin dimers are GTP-bound; they undergo GTP hydrolysis upon incorporation into MTs, resulting in a MT lattice that mainly consists of GDP-tubulin, and MT ends where GTP-tubulin prevails. In cells MT minus-ends are often embedded, whereas MT plus ends display stochastic cycles of polymerisation and depolymerisation, a behaviour called dynamic instability.
The α and β tubulin genes in mammals have been duplicated several times, giving rise to multiple highly homologous tubulin isotypes that are expressed in a tissue- and cell type-specific manner and that confer subtle properties to the MT they are part of. To simultaneously control isotype mRNA levels cells have a post-transcriptional mechanism in place, called tubulin autoregulation, in which imbalances in tubulin concentration at the protein level influence all the different tubulin-encoding mRNAs. Autoregulation was discovered in studies with MT-active agents (MTAs), compounds that bind tubulin on various sites and that deregulate MTs either by stabilising the network or by de-stabilising it. The best-known MT stabiliser is paclitaxel (PTX), or taxol. In contrast, colchicine and nocodazole depolymerise MTs. A key factor in autoregulation is the protein TTC5 which recognises nascent translating α and β tubulin polypeptides at the ribosome exit tunnel, and initiates mRNA degradation when tubulin levels are high. TTC5 is constitutively active in cells but its activity is low in normal conditions. Physiological functions for autoregulation remain poorly characterised.
While tubulin and MTs have been intensively examined from a mechanical perspective, roles for tubulin beyond MT assembly have been less well studied. We have previously established a system in which mammalian α and β tubulin are transiently co-expressed in equimolar amounts in cells. Here, we generated a Doxycycline (Dox)-inducible cell line to allow co-expression of α/β tubulin in a controlled manner. We examined MT behaviour and dynamics in cells overexpressing α/β tubulin, as well as the consequences of a mild tubulin overexpression. We find that overexpression generates surplus tubulin, despite the activation of tubulin autoregulation, as well as an excess of MTs. This perturbs MT behaviour and multiple processes in cells, including the cell cycle, which results in genome maintenance problems. Surprisingly, tubulin overexpression also induces mitochondrial dysfunction, skews the proteome by translation attenuation, and affects other stress responses. Conversely, amino acid and oxygen deprivation in normal cells suppress tubulin and the MT network.
Results
Characterisation of an inducible 293F cell line overexpressing a- and b-tubulin
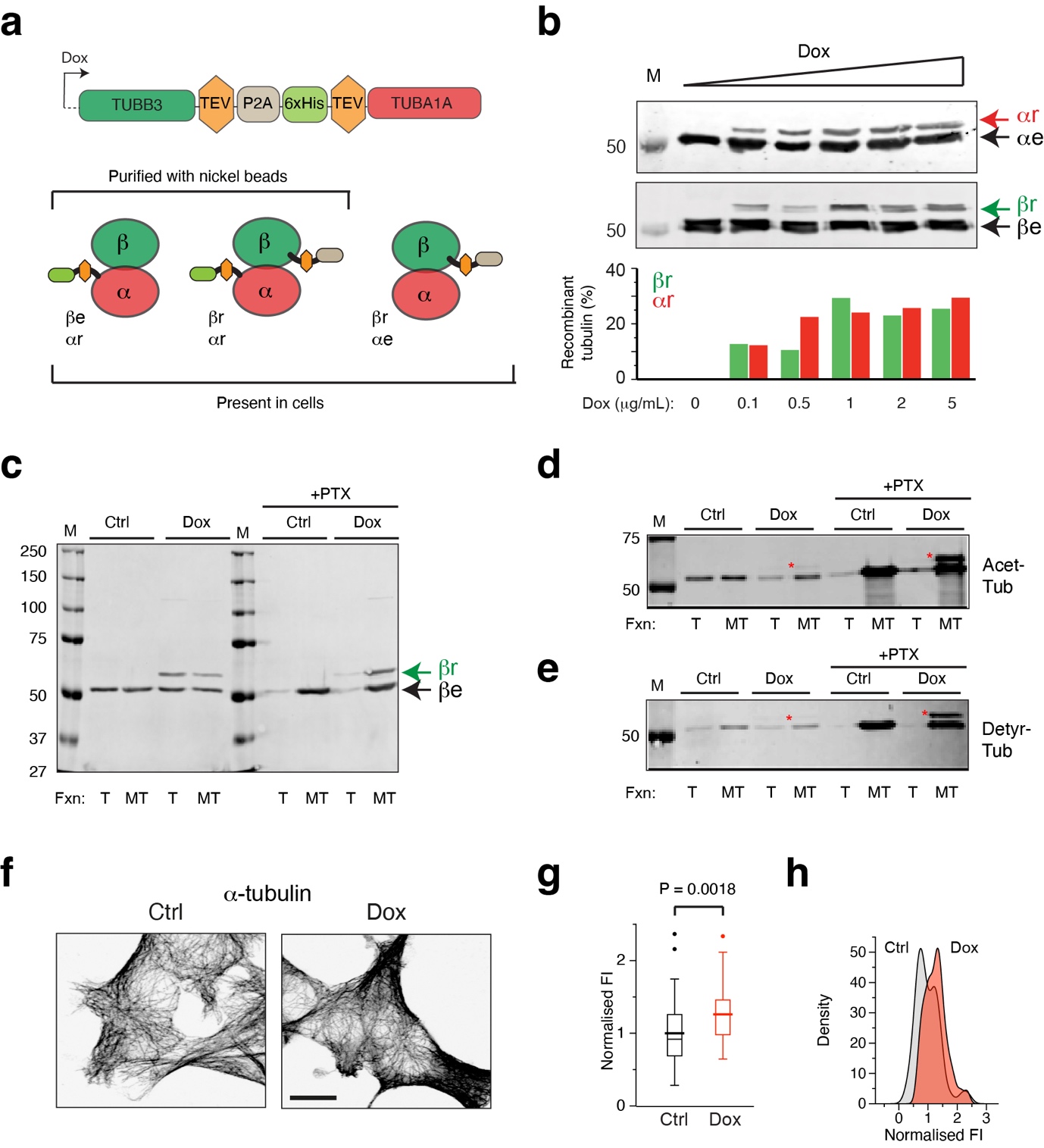
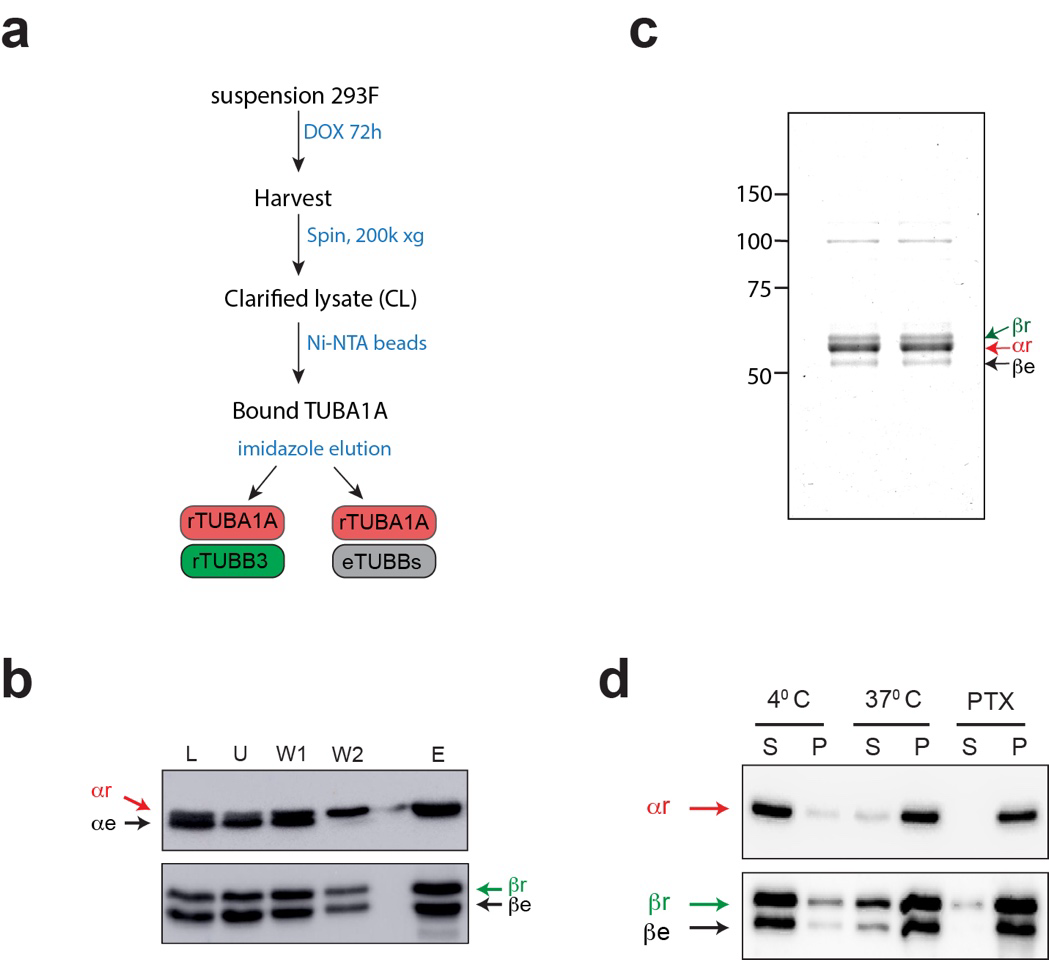
We used a “dual tubulin” strategy to produce novel human recombinant α and β tubulins from a single mRNA. Recombinant TUBB3 (rTUBB3) and TUBA1A (rTUBA1A), which can dimerise with each other and with endogenous tubulins, are distinguished on western blot and their levels with respect to endogenous tubulins can therefore be easily quantified.
The dual construct was placed downstream of a Tet-responsive element, and transfected into human 293F cells engineered to express a transgenic reverse-Tet transactivator (rTTA) gene. We generated multiple clones containing a stably integrated construct and selected one line that expressed rTUBB3 and rTUBA1A up to ~30% of endogenous tubulin levels. To examine functionality, we isolated recombinant tubulin using Ni-NTA affinity chromatography. We obtained purified rTUBA1A, dimerised with rTUBB3 as well as endogenous β-tubulin. We then performed MT polymerisation assays, at different temperatures and with or without PTX. In the presence of PTX more than 90% of all tubulin (recombinant and endogenous) was found in the MT pellet, whereas at 4°C hardly any tubulin polymerised, demonstrating that purified recombinant tubulin is assembly-competent.
To investigate recombinant tubulin properties in cells we fractioned cell lysates using high speed centrifugation. Recombinant tubulin was detected both in the soluble (tubulin-containing) and pellet (MT-containing) fractions. Addition of PTX for two hours prior to cell isolation greatly increased the amount of MTs, which also contained recombinant tubulin. Furthermore, two common post-translational modifications (PTMs) on alpha-tubulin in MTs, i.e. acetylation at the K40 residue and detyrosination (removal of the terminal Y residue), were observed on recombinant α-tubulin. We detected an increase of these PTMs on recombinant tubulin after pre-treatment of cells with PTX.
We next examined the interphase MT network in Dox-induced and control 293F cells using immunofluorescence microscopy. Fluorescence intensity of the secondary antibody was taken as a readout of MT density. We observed a 1.3-fold higher fluorescence intensity in Dox-induced cells compared to control cells. The whole cell population contained excess MTs, indicating that Doxycycline induces an excess of tubulins and MTs in all cells. This situation is unique compared to cells treated with high concentrations of MTAs, which either lead to an increase in MT mass with concomitant loss of soluble tubulins (e.g. PTX treatment), or to an increase in soluble tubulins and a loss of MTs (e.g. nocodazole or colchicine treatments).
Tubulin overexpression affects dynamic MT behaviour
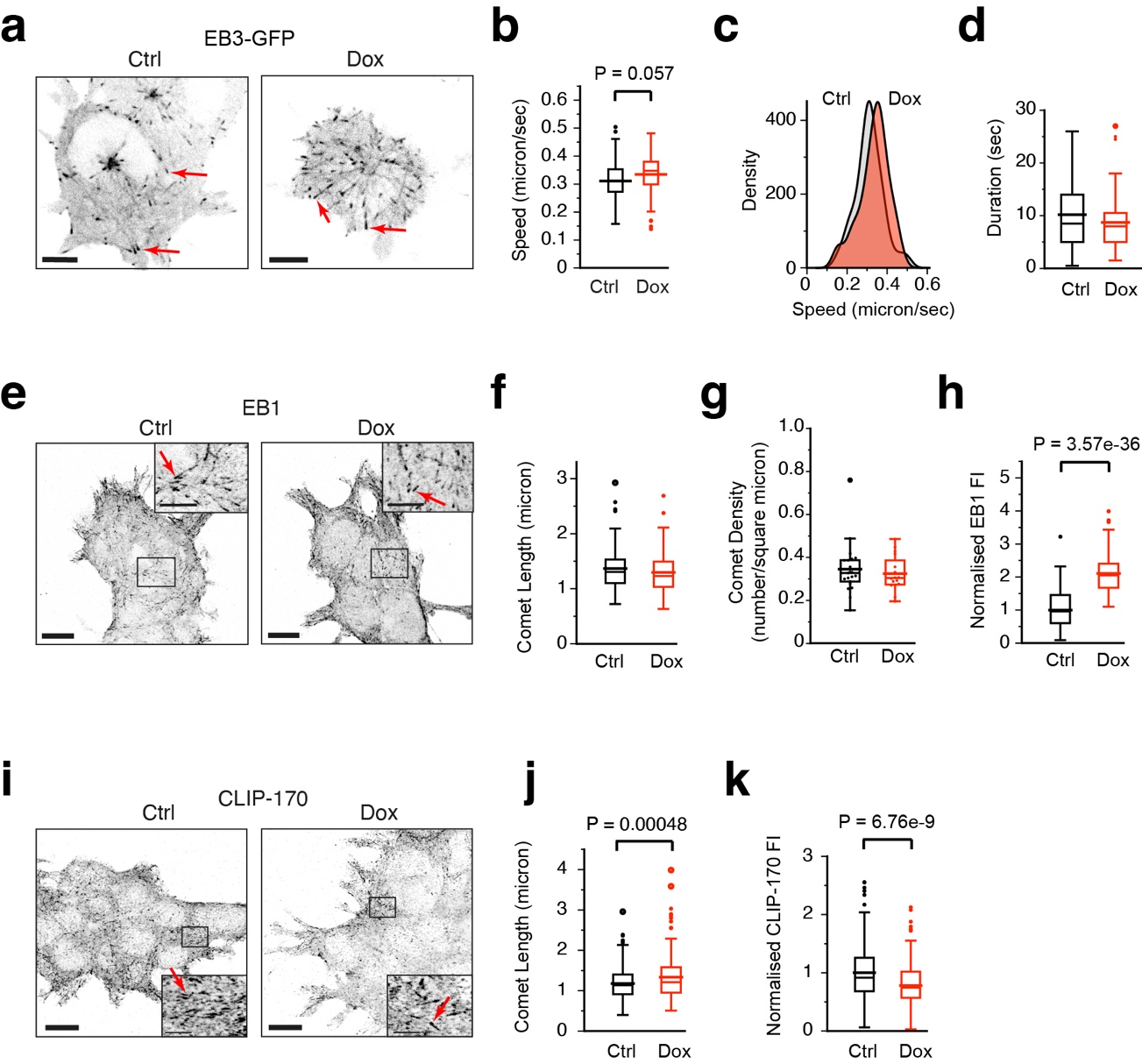
To assess the effect of dual tubulin overexpression on MT dynamics we transfected control and Dox-induced cells with a cDNA encoding the MT plus-end marker EB3-GFP. We acquired time-lapse videos of EB3-GFP 24 hr after transfection and examined MT growth rates as well as growth duration (the inverse of catastrophe frequency). We detected a mild increase in MT growth rates in Dox-treated cells, which was observed in all of the growing MTs examined, consistent with an increase in soluble tubulin levels in all Dox-induced cells. Surprisingly, a slight decrease in growth duration was detected, indicating that although MTs grow faster after Dox addition the rate of catastrophe is slightly higher.
We next examined the localisation of the +TIPs EB1 and CLIP-170. Immunofluorescence (IF) staining for EB1 did not reveal differences with respect to comet length or overall number of plus-ends per unit area. However, the fluorescence intensity of EB1 at the MT end was approximately 2-fold higher in Dox-induced cells, indicating that twice as many EB1 proteins are bound per MT end in Dox-induced cells. By contrast, IF stainings for CLIP-170 revealed an increased CLIP-170 comet length in Dox-induced cells, whereas fluorescence intensity at the MT end was decreased.
Mitotic defects in 293F cells overexpressing tubulin
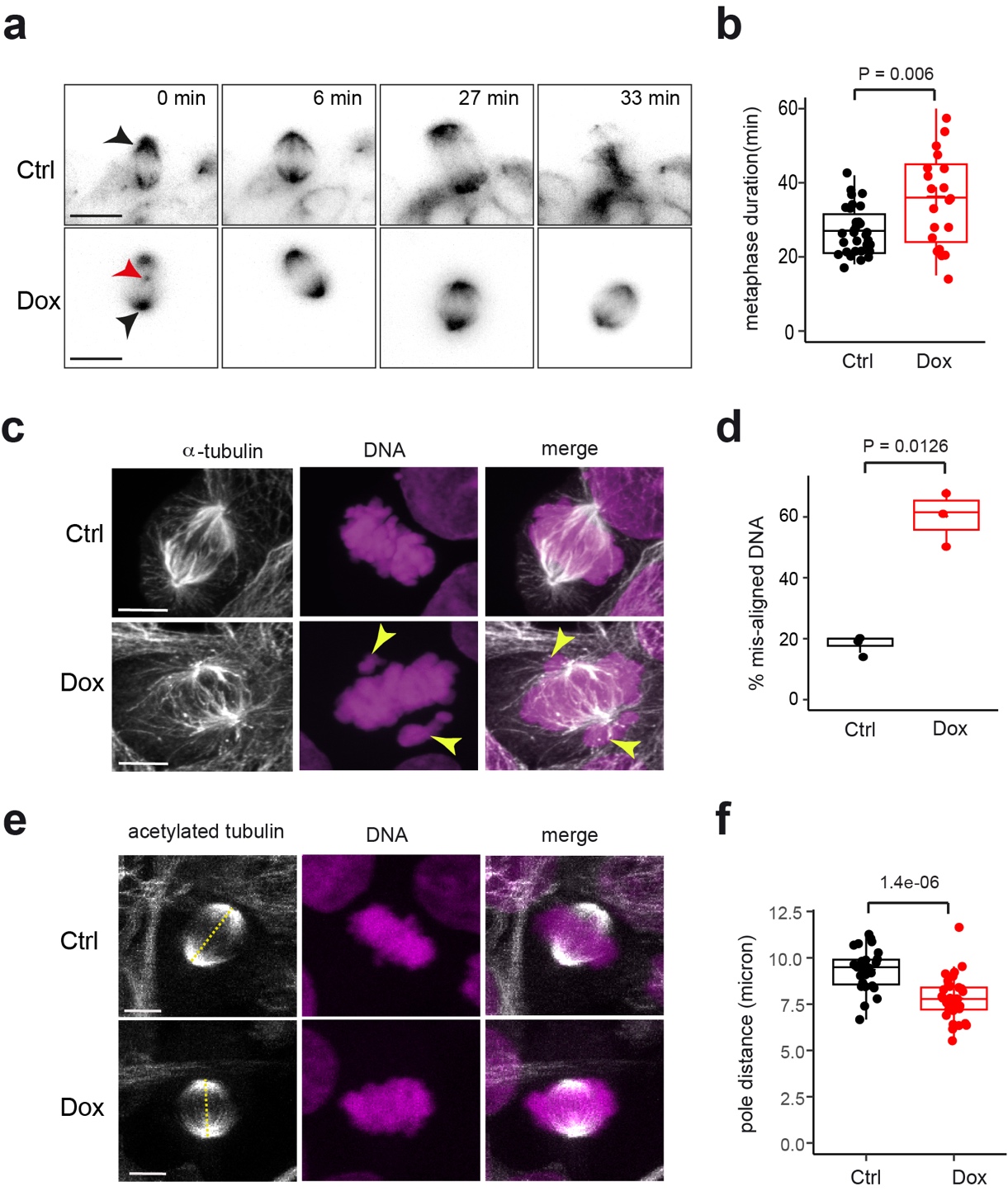
Light sheet fluorescence microscopy (LSFM) revealed that the average time taken to complete metaphase was higher after Dox-induction, indicating that the spindle assembly checkpoint (SAC) is activated in Dox-induced cells to delay anaphase onset. We also observed “extra-mitotic” centrioles at the onset of mitosis after Dox-treatment, indicative of mitotic arrest and aneuploidy.
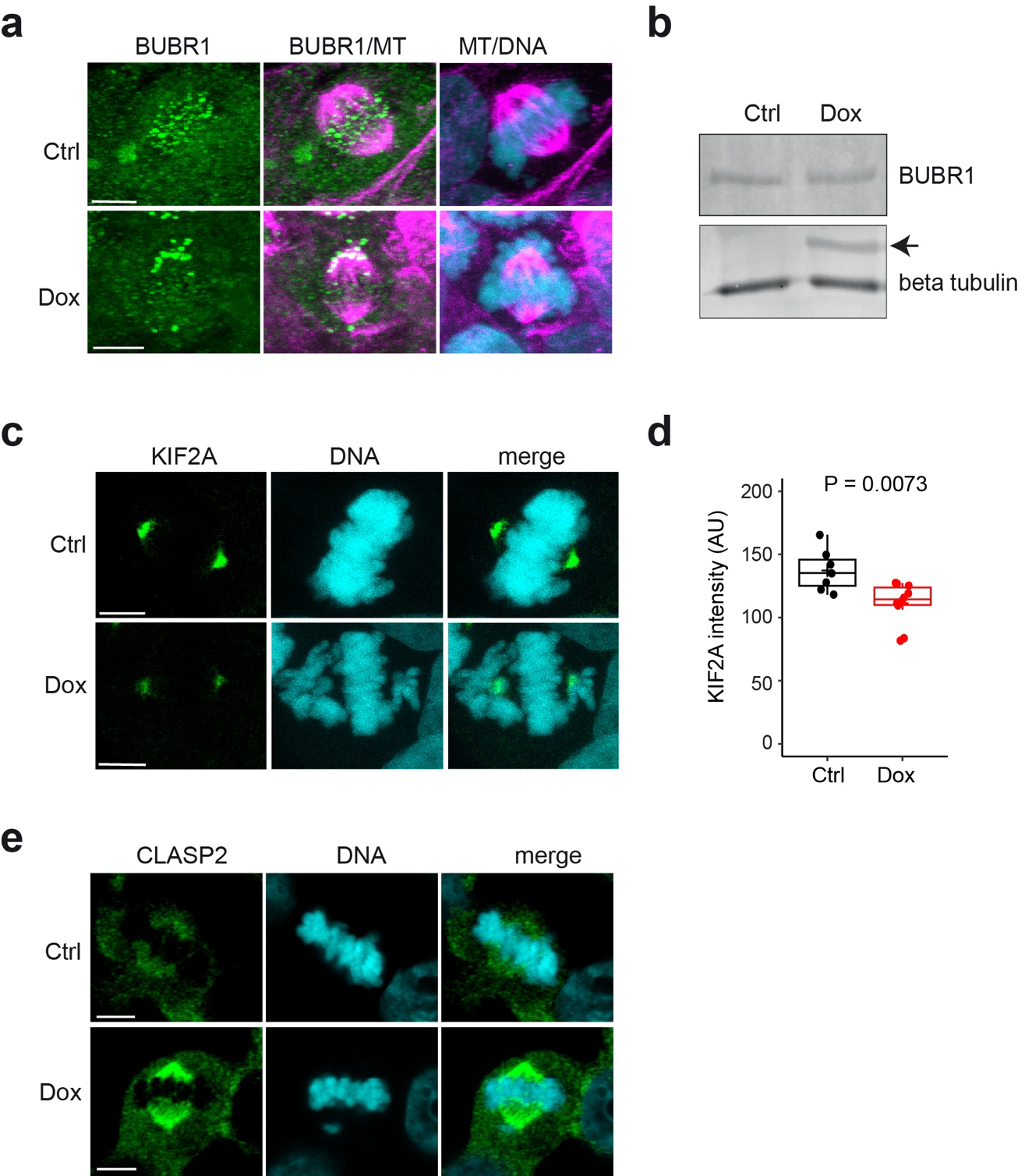
When we examined mitotic structures in fixed cells, we observed increased chromosome mis-alignment at the metaphase plate, indicating that chromosome segregation was impaired in Dox-induced cells. We found aberrant BUBR1 accumulation at the metaphase plate in Dox-treated cells. Western blotting for BUBR1 in cell lysates did not show a change in protein level after Dox-induction. These data suggest that mild tubulin-overexpression engages the SAC and delays anaphase onset.
Finally, we found that metaphase spindles had an overall reduced length after Dox-addition. Furthermore, the mitotic defects were accompanied by mis-localisation of spindle-associated proteins, as the levels of the kinesin KIF2A, which localizes at spindle poles where it acts to depolymerize MTs, were reduced in Dox-induced cells, while CLASP2, which is mainly localised to kinetochore MTs, was increased after Dox-induction.
Tubulin overexpression deregulates the cell cycle and affects genome integrity
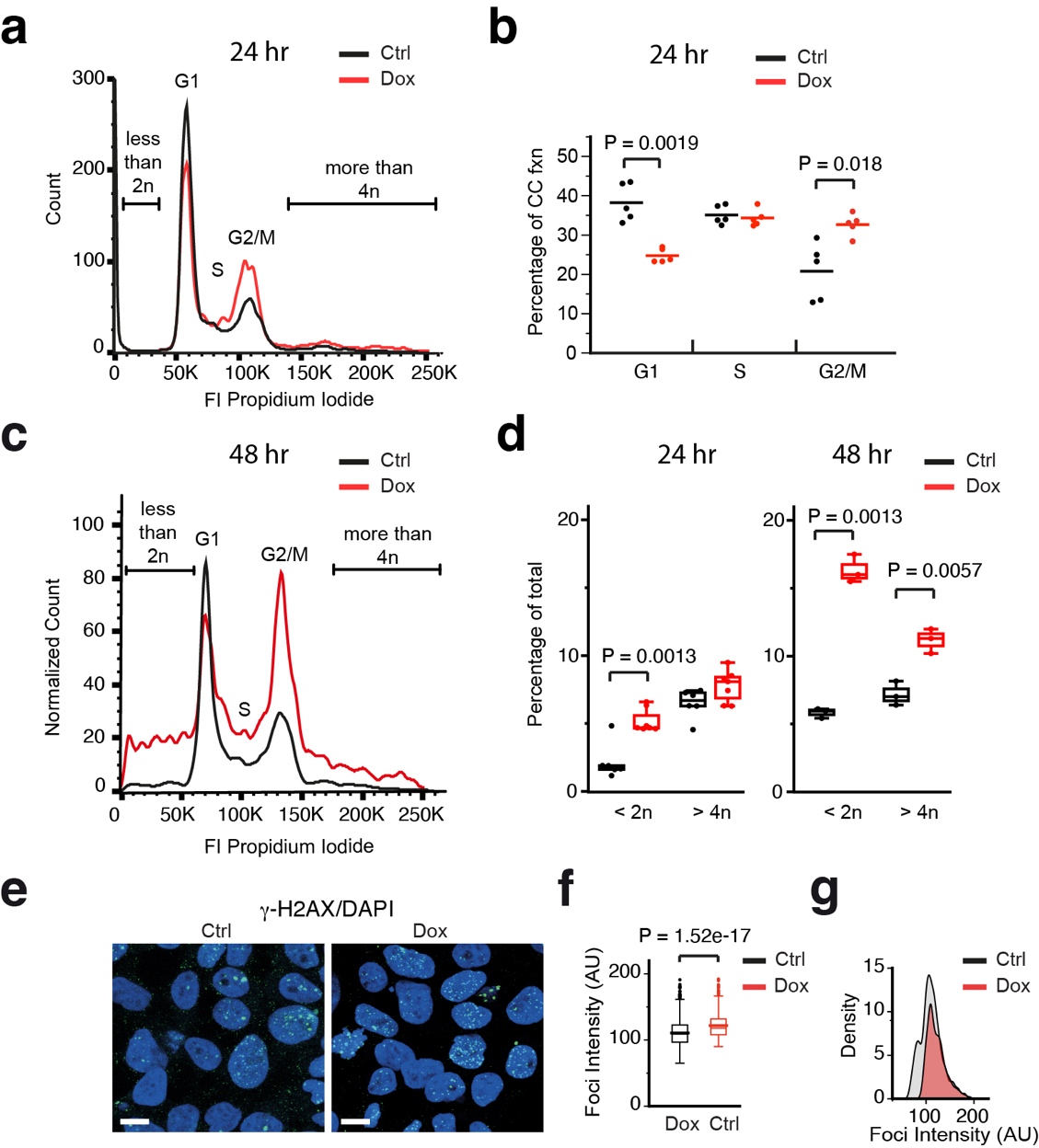
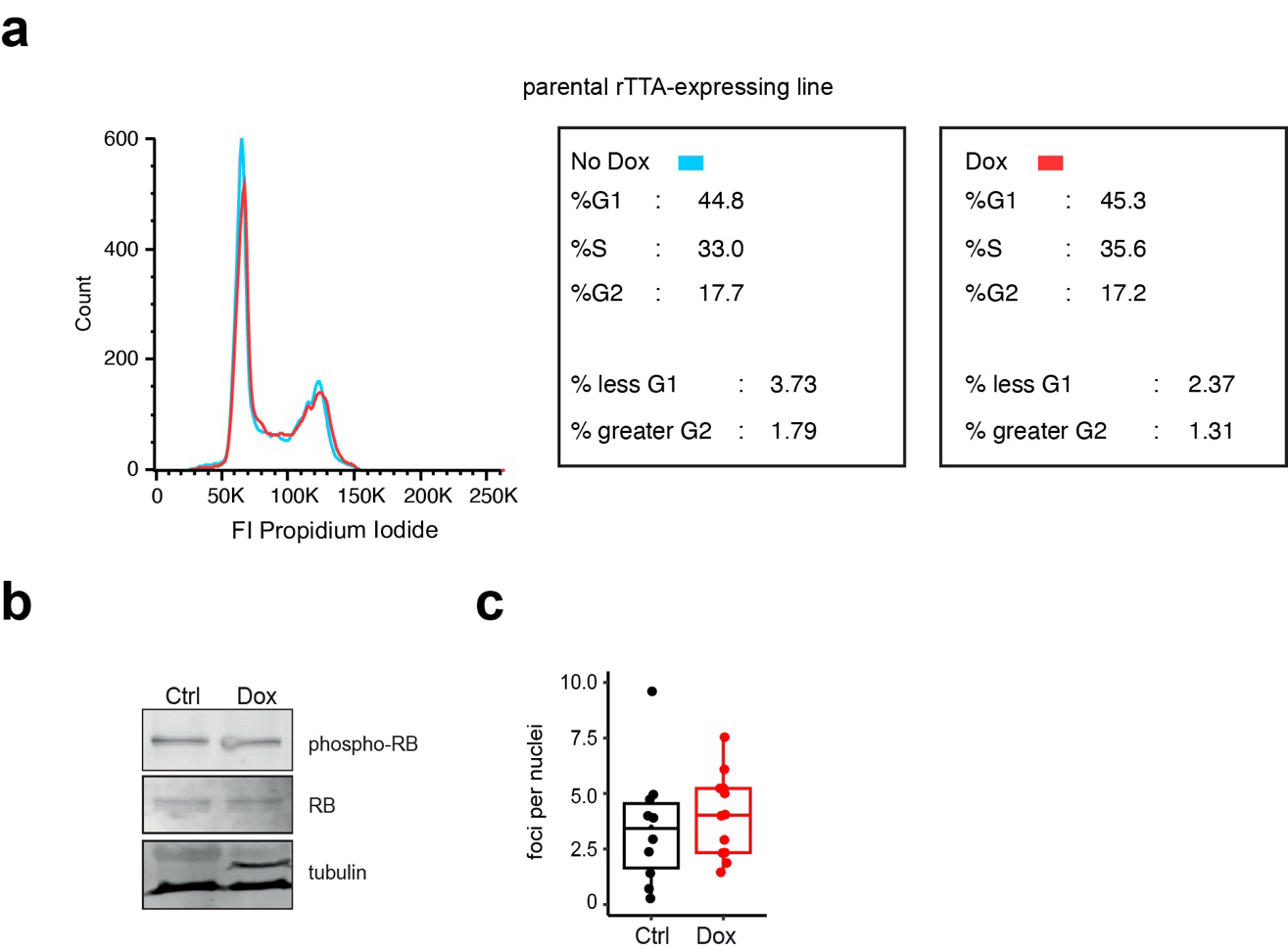
We examined the cell cycle in tubulin overexpressing cells using flow cytometry. Cells were fixed after 24 or 48 hours of Dox-induction, and stained with propidium iodide (PI) to investigate DNA content. No effect of Doxycycline was observed in the parental Tet-expressing cell line after 24 hr Dox-induction, indicating that neither Doxycycline itself nor rTTA expression influences the cell cycle. By contrast, we detected a decrease in the G1 population and an increase in the G2/M fraction after 24 hour of tubulin overexpression. Surprisingly, despite the reduction in the G1 fraction, similar S-phase cell populations were detected in the control and dox-induced cells. After 48 hr of Dox-induction all fractions (G1, S and G2/M) were perturbed. Moreover, DNA content was highly aberrant after 48 hr, with large proportions of cells containing < 2n or > 4n chromosomes.
The level of phospho-RB was similar in Dox-induced and control 293F cells, despite a decreased number of G1 cells in the Dox-induced population. We therefore hypothesised that tubulin overexpression leads to a premature release of the G1/S restriction point. Using an antibody against γH2A-X, which detects the phosphorylated form of Histone H2A at double strand break (DSB) lesions, we detected no significant increase in the number of γH2A-X-positive foci observed per nucleus, but did observe a higher intensity of γH2A-X signal within foci after Dox-induction, suggesting that tubulin overexpression results in an accumulation of DSB lesions due to slower repair.
RNA-sequencing reveals autoregulation in tubulin overexpressing cells
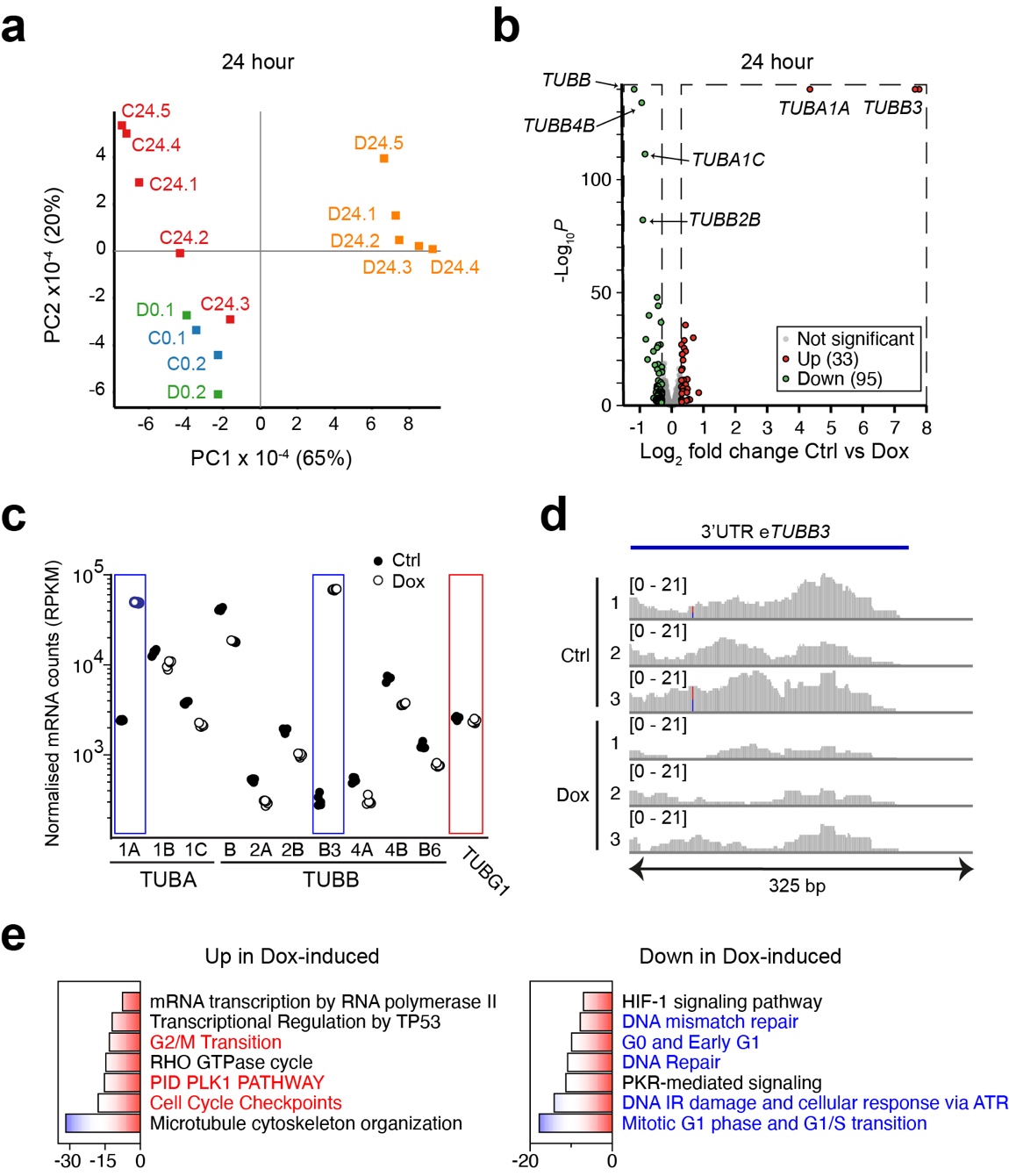
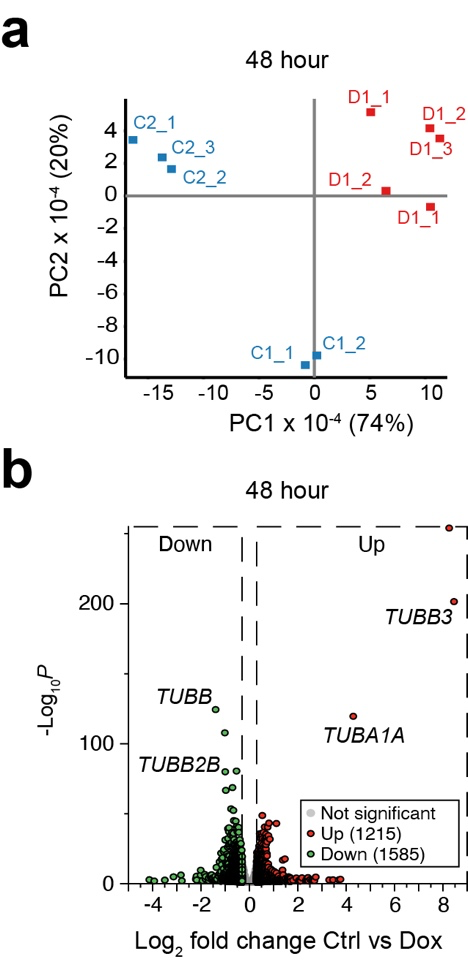
To analyse the genome-wide consequences of tubulin overexpression, we performed RNA-sequencing (RNA-seq) on control and Dox-induced cells. Samples were either collected at the start of the experiment (0 hr), or after 24 or 48 hr. Principal component analysis (PCA) of the 0 and 24 hr data showed that the 24 hr Dox-induced samples clustered together and were separated from the 24 hr control and the 0 hr samples.
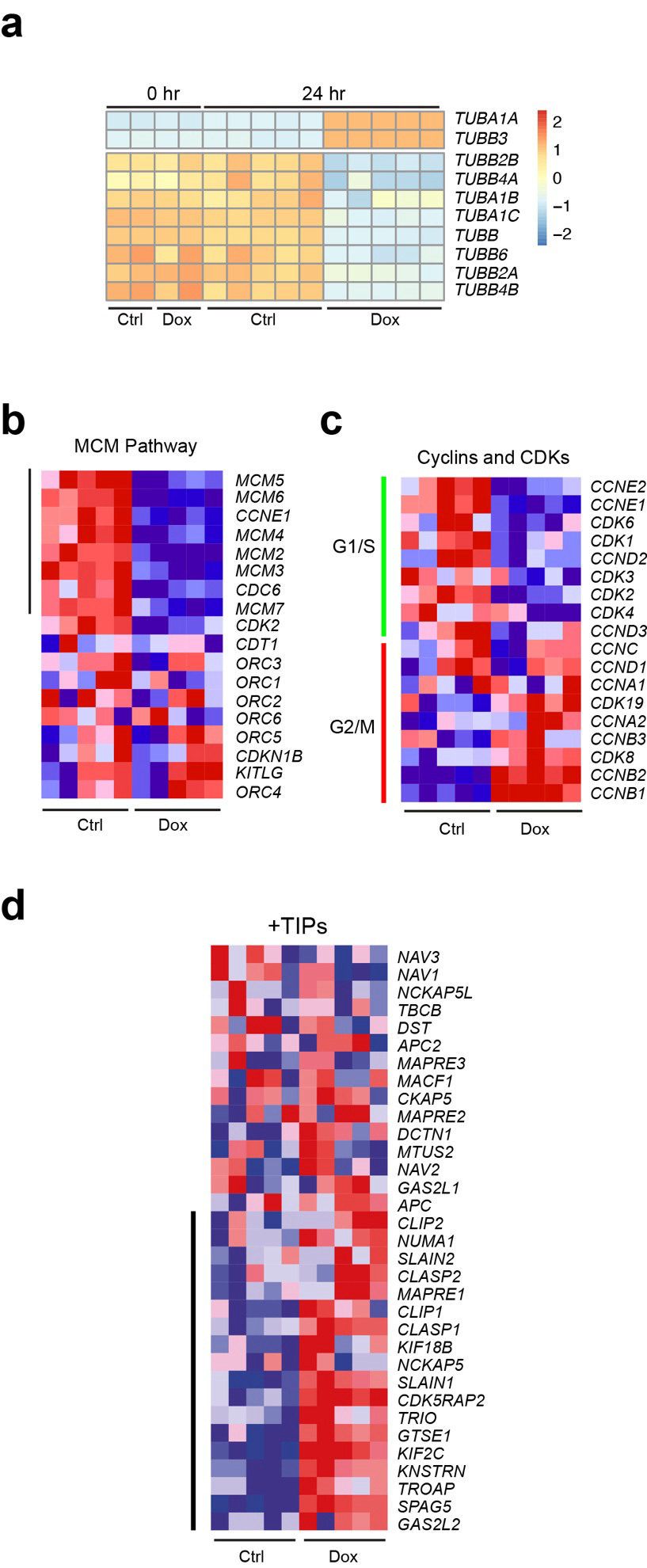
To examine differentially expressed genes (DEGs) we performed a DESeq2 analysis. Not many mRNAs were significantly deregulated after 24 hr Dox-induction (33 mRNAs up and 95 mRNAs down). By contrast, many more DEGs were observed after 48 hr of Dox-induction (1215 mRNAs up and 1585 down). Notably, both after 24 hr and 48 hr tubulin-encoding mRNAs were among the most affected DEGs. To assess tubulin expression in more detail we extracted the reads from tubulin-encoding mRNAs after 24 hr Dox-induction. We observed upregulation of recombinant TUBA1A and TUBB3 in the Dox-induced samples, and downregulation of endogenous tubulin-encoding mRNAs. Importantly, the extent of downregulation was similar for all isotypes, strongly suggesting that autoregulation is the mechanism by which tubulin mRNAs are curbed and not transcriptional control, which will not act on all tubulin isotype genes.
RNA-sequencing reveals an altered stress response in tubulin overexpressing cells
We next analysed the RNA-seq data for effects beyond autoregulation. We performed both a metascape analysis on DEGs, and a Gene Set Enrichment Analysis (GSEA) on the full RNA-Seq dataset. Metascape analysis revealed downregulation of G1-S- and DNA replication-related pathways in Dox-induced cells, and upregulation of G2-related pathways, including PLK1, and of mitotic spindle components. GSEA confirmed the Metascape results, as the G1-related MCM pathway was down in Dox-induced cells.
In addition to cell cycle defects, the Metascape analysis suggested that tubulin overexpression affected HIF-1- and PKR-mediated signaling. Hypoxia-inducible factor 1 (HIF-1) is a transcription factor that regulates the cellular response to oxygen deprivation, or hypoxia. Protein kinase double-stranded RNA-dependent pathway (PKR), which is induced by various stress mechanisms including viral infections, acts in the integrated stress response (ISR) and mainly halts protein translation in response to stress by phosphorylating EIF2α. The fact that HIF-1 and PKR signaling were dampened in Dox-induced cells was surprising and suggested that tubulin overexpression somehow hampers these responses.
Tubulin overexpression causes mitochondrial and proteostasis defects
Since RNA-Seq does not necessarily detect changes at the protein level, we decided to perform proteomic profiling of Dox-induced 293F cells. We cultured Dox-induced and control cells for 24 hr under 2.5% oxygen conditions (hypoxia) or 20% oxygen conditions (normoxia) and analysed cellular proteomes using mass spectrometry. We detected 8224 proteins in these experiments, of which 3128 passed an ANOVA-based significance threshold.
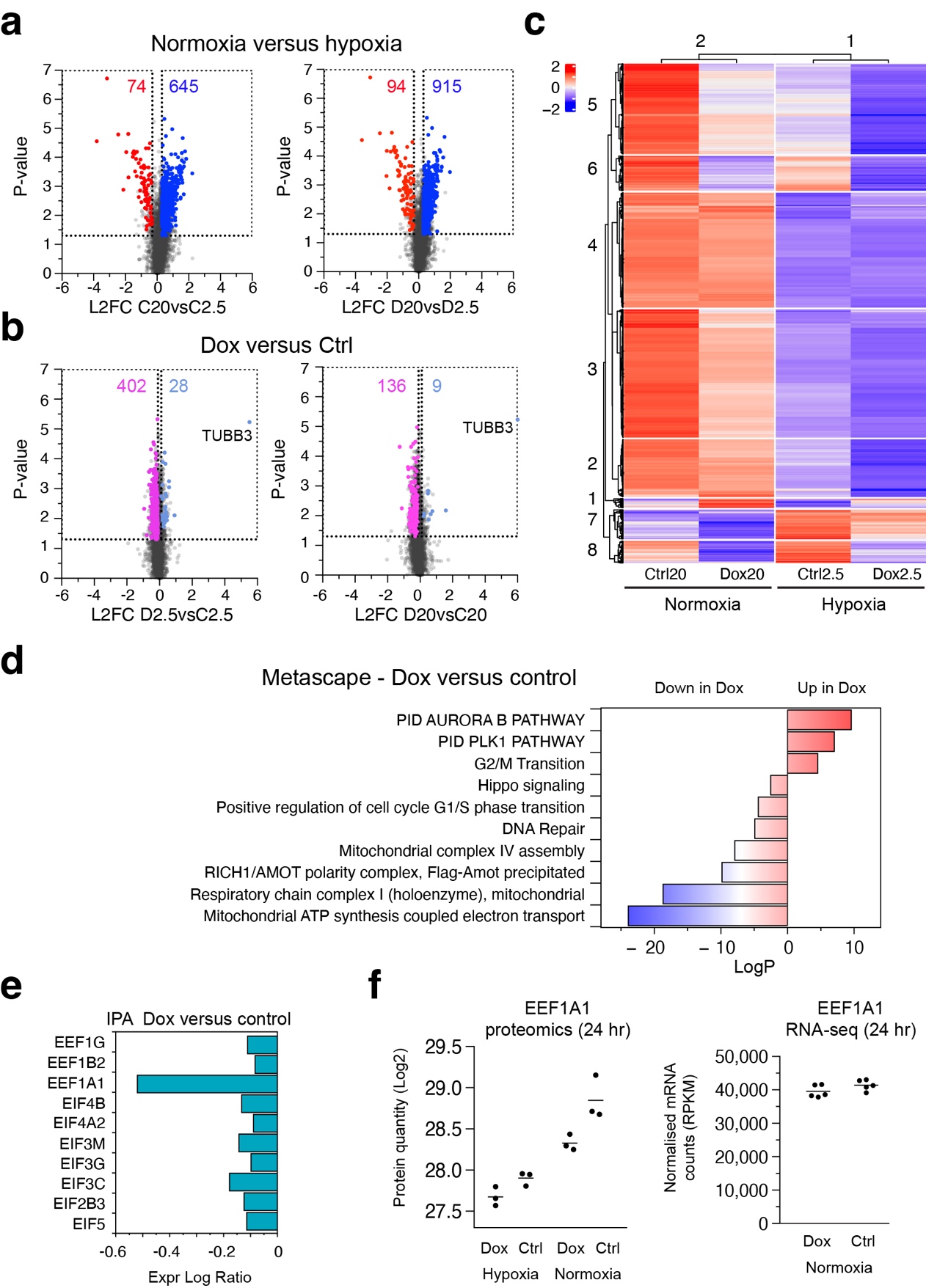
Volcano plots of the proteomes in different conditions revealed many more proteins upregulated in normoxia compared to hypoxia, both in control and Dox-induced cells. This result is expected since hypoxia can result in general translation inhibition due to the ISR. Surprisingly, we also observed skewing when we compared Dox-induced and control cells, with many more proteins down in the Dox-induced cells, both in hypoxia and normoxia. Although the skewing was less pronounced compared to hypoxia versus normoxia, the data demonstrate that tubulin overexpression causes proteostasis defects.
A heatmap representation of a K-means clustering analysis visually confirmed the reduced protein levels in hypoxic conditions and after dox-treatment. Only in protein cluster 7, which consisted of 195 proteins, did we observe increased protein levels in hypoxic conditions. This cluster contained the HIF1 response pathway as the top term in a Metascape analysis. Thus, both control and Dox-induced cells activate HIF1 in hypoxia. However, HIF1 targets are present at lower levels in Dox-induced cells. The HIF1 response is therefore dampened by tubulin overexpression.
To uncover the mechanism by which tubulin overexpression causes proteostasis defects we focussed on the Dox-induced versus control samples. Metascape analysis revealed upregulation of terms associated with mitosis and G2/M transition, including the PLK1 pathway, in the Dox-induced cells, whereas G1/S-specific, and DNA repair terms were down. Surprisingly, a number of terms related to mitochondria were found to be downregulated in the Dox-induced cells, including electron transport, respiration, and mitochondrial complex I and IV assembly.
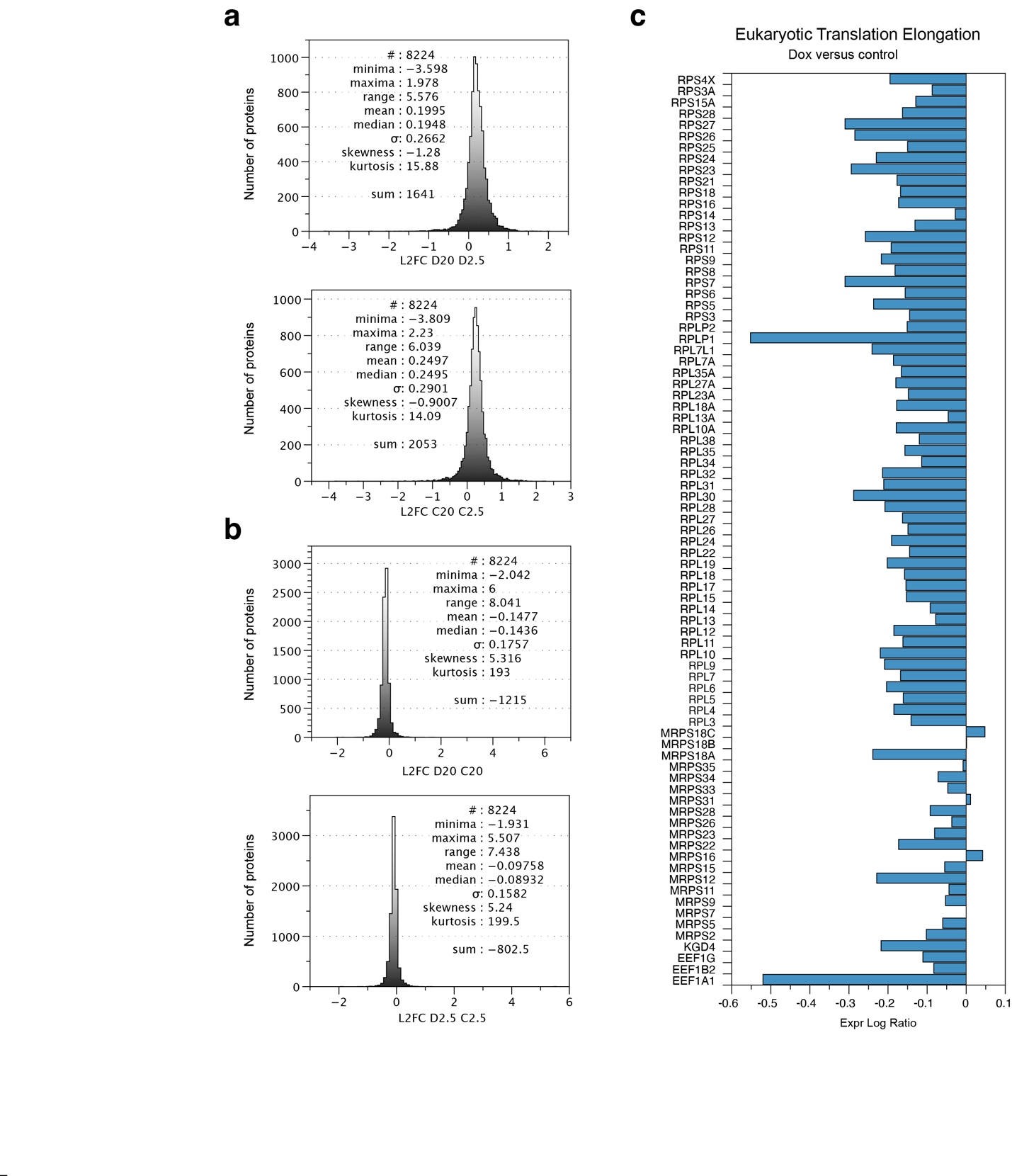
Proteostasis defects can be caused by alterations in translation efficiency. To analyse possible effects on translation after tubulin overexpression we performed IPA on an extended protein dataset. Strikingly, top hits in IPA revealed terms involving translation, including Major pathway of rRNA processing in the nucleolus and cytosol, Eukaryotic Translation Initiation, Eukaryotic Translation Elongation, and Eukaryotic Translation Termination. Virtually all proteins in the pathways were downregulated in the Dox-induced cells.
Transient mitochondrial stress causes translation attenuation due to defects in respiration and ensuing lowered levels of the translation elongation factor EEF1A1. On the other hand, prolonged mitochondrial stress eventually results in the ISR and hampers translation initiation. To distinguish between these two, we examined the levels of significantly deregulated initiation and elongation factors in the Dox-induced proteome, and found that EEF1A1 was most affected. Notably, EEF1A1 was downregulated at the protein level in Dox-induced cells, both in normoxia and hypoxia, but not at the mRNA level, indicating that EEF1A1 itself is translationally regulated.
Characterisation of the tubulome of 293F cells
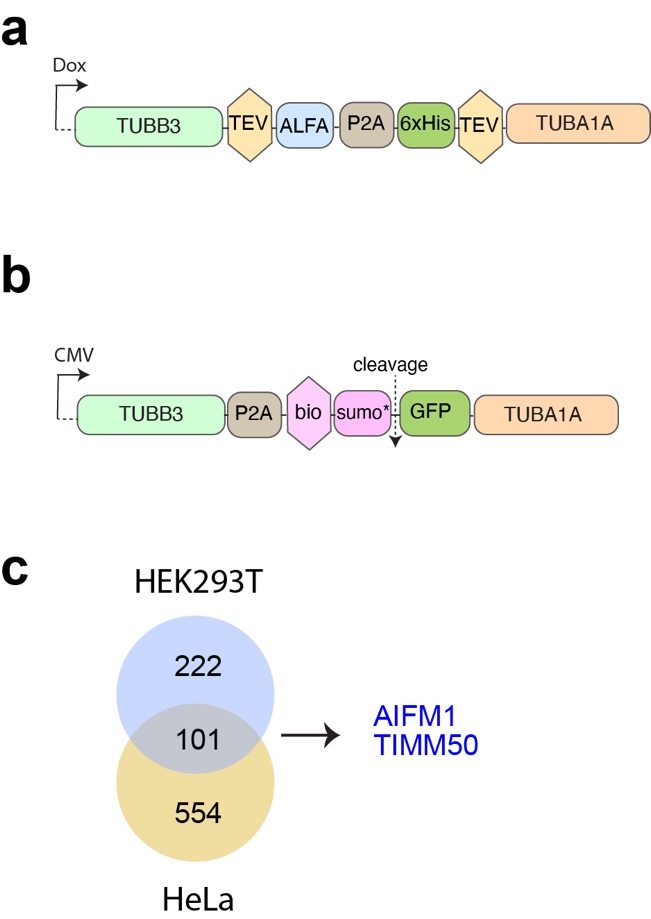
To uncover tubulin-mediated mechanisms underlying proteostasis defects we investigated the tubulin interactome, or “tubulome”, of 293F cells. We generated a stable 293F cell line in which β-tubulin was tagged at its C-terminus with the ALFA-tag. Using beads coupled to ALFA-tag antibodies, we affinity-purified recombinant tubulin and tubulin-associated proteins (TAPs) from cell lysates of tubulin overexpressing cells, using non-induced cells as controls.
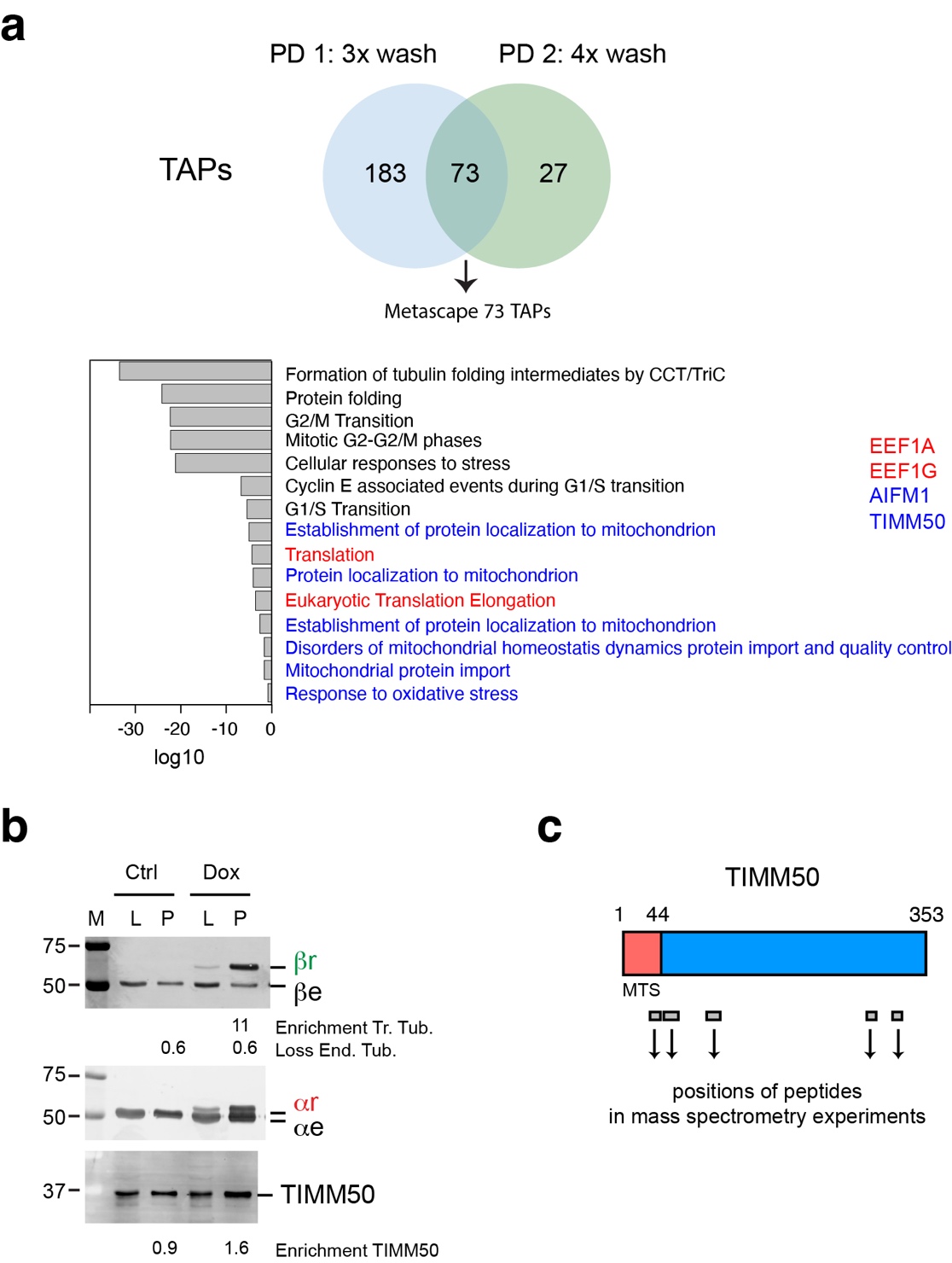
We observed 73 TAPs coming down in both pull downs. Metascape analysis of these common TAPs revealed tubulin folding as the strongest term. Since chaperones and tubulin folding factors are well known TAPs, these results validate our tubulin interaction approach in stably expressing 293F cells. Importantly, Metascape analysis suggested that common TAPs are involved in the very processes affected by tubulin overexpression in 293F cells. For example, we detected TAPs involved in G2/M and G1/S transition, and in DNA damage checkpoints. In addition, five terms involving mitochondrial homeostasis were observed, including mitochondrial import and quality control, and two terms involving translation.
The mitochondrial terms consistently contained the proteins AIFM1 and TIMM50. AIFM1 (Apoptosis Inducing Factor Mitochondria Associated 1) is a mitochondrial flavoprotein that was first associated with apoptosis, but was later found to be required for the normal expression of major respiratory chain complexes. TIMM50 is involved in the import of nuclear-encoded mitochondrial proteins into mitochondria. We validated the interaction between TIMM50 and recombinant tubulin on western blot using antibodies against TIMM50 and tubulins.
When we mapped the peptides detected by mass spectrometry to TIMM50, we found that one of them was located within the MTS (mitochondrial targeting sequence). These results suggest that tubulin interacts with the precursor form of TIMM50. Based on our results we propose that excess tubulin binds and sequesters TIMM50, thereby hampering protein import into mitochondria, specifically of proteins involved in electron transport and respiration.
Regulation of the MT cytoskeleton during cellular stress
We have shown that overexpression of tubulin affects the stress response of cells. We then asked whether the converse also occurs, namely that stress responses affect tubulin levels. We found evidence for this hypothesis in a landmark study which showed that autoregulation is active in many experimental conditions in cells and even in vivo. As part of a dataset verification, the authors examined glutamine (Gln) deprivation in U2OS cells and observed a decrease in tubulin at the mRNA and protein level, as well as a reduction in MTs, after 24 hr deprivation.
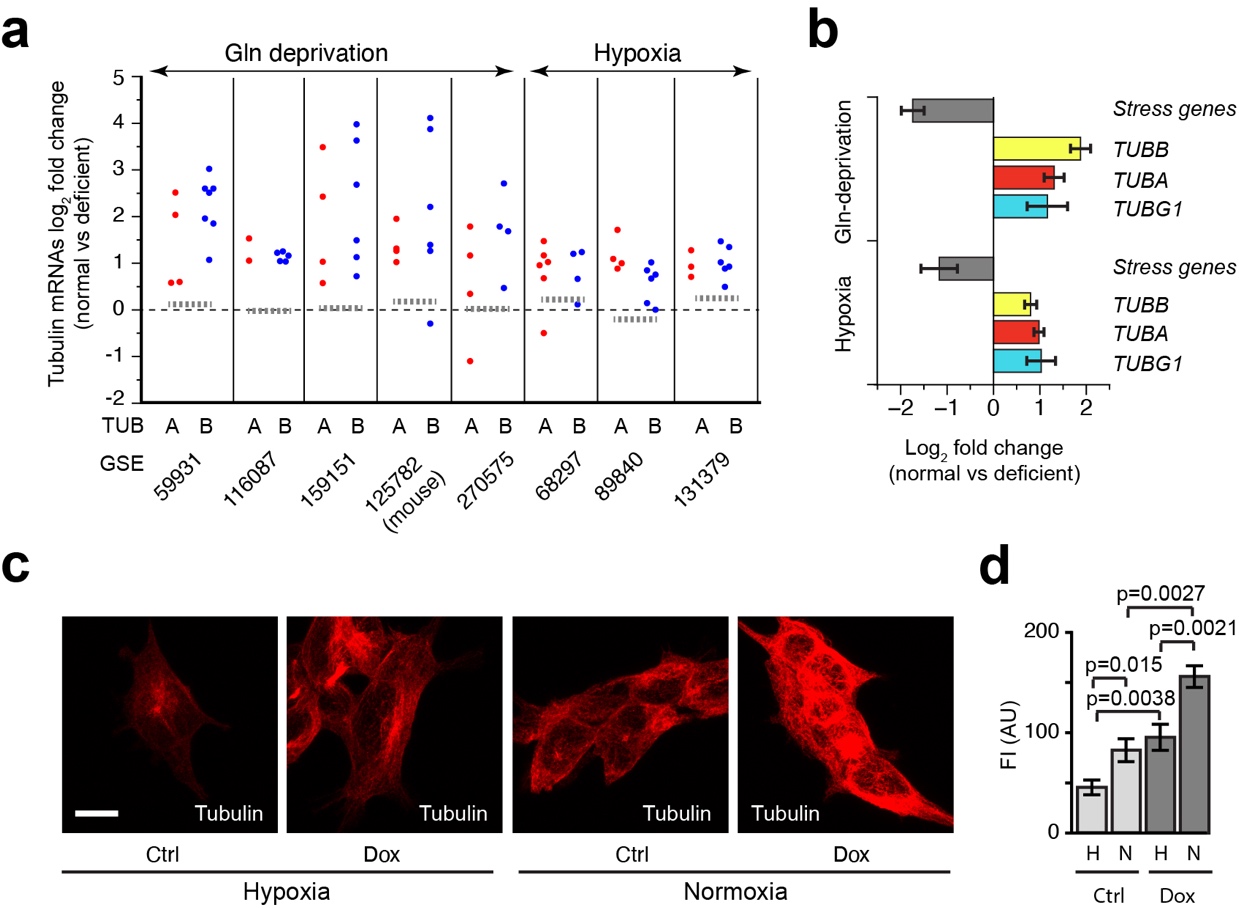
To address this, we retrieved multiple transcriptomic datasets in which either hypoxia or Gln deprivation were studied. We then extracted log2-fold changes in expression of tubulin-encoding mRNAs and selected validated stress-related mRNAs as controls. We observed downregulation of virtually all the alpha- and beta-tubulin-encoding mRNAs, both after Gln deprivation and hypoxia. By contrast, stress-related genes were upregulated, as expected. Since the tubulin-encoding mRNAs are regulated in the same manner, the results indicate that autoregulation is activated concomitant with the response to amino acid or oxygen deprivation. Interestingly, TUBG1 was downregulated together with tubulin-encoding mRNAs, as would be expected if autoregulation is activated.
To assess whether stress affected MTs in our system, we cultured control and Dox-induced 293F cells for 24 hr under hypoxic or normoxic conditions, and analysed the network using immunofluorescence microscopy and tubulin antibodies. We found that hypoxia significantly reduced cellular MT density in control cells, consistent with the reduction in tubulin- and TUBG1-encoding mRNAs seen in the hypoxic RNA-seq datasets. Since a decrease in MTs has also been observed in glutamine-deprived U2OS cells, the data suggest that the MT cytoskeleton is dampened in at least two different stress responses. Consistent with the data presented in earlier experiments, the Dox-induced cells displayed a more abundant MT network compared to controls. Interestingly, hypoxia treatment did decrease the MT network in tubulin overexpressing cells, but not to the extent observed in control cells.
Discussion
In this work, we utilised a Doxycycline-inducible P2A-based expression system to produce recombinant α/β-tubulin subunits in equimolar amounts. We show that recombinant tubulins are functional in vitro and in cells, and that mild overexpression leads to an excess of soluble tubulin and MTs. The overabundance of MTs indicates that they serve as a storage reservoir for excess tubulin. Although this is logical it does raise the question why, in contrast to tubulin, cells do not sense excess MTs despite the presence of a variety of MAPs, including MT depolymerases. A main reason could be that deviations in MT structure and density are a normal occurrence in cells, serving particular functions.
Our RNA-seq analysis showed that a mild overexpression of recombinant tubulins is sufficient to engage autoregulation. Physiological functions of this post-transcriptional regulatory mechanism have not yet been investigated in detail. An early hypothesis was that by controlling tubulin levels autoregulation prevented spurious MT nucleation in cells. In agreement with this view, we propose that aberrant branched MT nucleation might underlie the mitotic phenotype of Dox-induced cells. Moreover, autoregulation may help to prevent the formation of excess MTs in cells, which seem to lack a MT-based surplus sensing mechanism. A corollary of this is seen in stress situations like Gln-deprivation and hypoxia, where transcriptomic dataset analysis suggests that autoregulation is activated and that this serves to downregulate MTs. Indeed, immunofluorescence microscopy studies show that this correlates with a diminished MT network in hypoxic 293F cells, and, as shown earlier, with a reduction in MTs in Gln-deprived U2OS cells. Autoregulation therefore appears to contribute to the downregulation of the MT network in stress situations.
We found that tubulin overexpression induces mitochondrial dysfunction. We propose that excess tubulin sequesters TIMM50 and AIFM1 resulting in reduced mitochondrial import. Indeed, constituents of the mitochondrial electron transport chain and oxidative phosphorylation, shown to be transported by TIMM50, were downregulated in tubulin overexpressing cells at the protein but not the mRNA level. Our mass spectrometry-based experiments furthermore show that tubulin overexpression results in proteostasis defects and suggest that this occurs by translation attenuation. Whereas the ISR inhibits general translation by phosphorylating EIF2α, acting on translation initiation, transient mitochondrial stress affects elongation by lowering the level of EEF1A1. Strikingly, we found that EEF1A1 protein levels are down in tubulin overexpressing cells. We therefore hypothesise that excess tubulin sequesters AIFM1 and TIMM50, resulting in a lowered mitochondrial protein import and stress, and that this in turn reduces EEF1A1, attenuates translation, and skews the proteome in Dox-induced cells. As we observed an interaction between tubulin and two translation elongation factors (EEF1A1 and EEF1G) in our ALFA-tag pull downs, we presume that tubulin overexpression can directly influence translation elongation.
TTC5 simultaneously binds the nascent translating tubulin polypeptide and the ribosome itself, and then recruits SCAPER and the CCR4-NOT deadenylase to degrade tubulin mRNAs. Studies with protein synthesis inhibitors revealed that TTC5 depends on ongoing translation to initiate autoregulation. In normal cells, translation is rapid and TTC5 therefore has a short time window to interact with the first amino acids of the tubulin peptide before they fold up. This explains, at least partly, why TTC5 activity is low in normal conditions. We provide evidence here that surplus tubulin slows general translation, increasing the time window for a TTC5-nascent tubulin interaction. TTC5 in turn relays specificity for tubulin mRNA degradation, and, as recently discovered, for TUBG1 mRNA degradation.
Sequestering of MAPs and TAPs is an overarching explanation for multiple phenomena observed in this report, with mitochondrial stress serving as a prime example. The premature G1/S release and ensuing replication stress observed in tubulin overexpressing cells could also be caused by the sequestering of an important protein involved in the G1/S checkpoint. Furthermore, HIF1 might be sequestered by surplus MTs in Dox-induced cells, preventing a proper hypoxia response. Altogether our data suggest that tubulin has multiple functions beyond the assembly of MTs. Here, we reveal a role in the three main cell cycle checkpoints, in cellular stress responses, and in translation. Our data explain why tubulin, a major cytoskeletal protein and among the more abundantly expressed proteins in a cell, needs to be exquisitely controlled to maintain a healthy genome and proteome.
References
Brouhard GJ, Rice LM. Microtubule dynamics: an interplay of biochemistry and mechanics. Nat Rev Mol Cell Biol. 2018;19:451-463. doi: 10.1038/s41580-018-0009-0
Desai A, Mitchison TJ. Microtubule polymerization dynamics. Annu Rev Cell Dev Biol. 1997;13:83-117. doi: 10.1146/annurev.cellbio.13.1.83
Goodson HV, Jonasson EM. Microtubules and Microtubule-Associated Proteins. Cold Spring Harb Perspect Biol. 2018;10(9):a022043. doi: 10.1101/cshperspect.a022043
Breuss M, Keays DA. Microtubules and neurodevelopmental disease: the movers and the makers. Adv Exp Med Biol. 2014;800:75-96. doi: 10.1007/978-3-642-55195-6_3
Cleveland DW. Autoregulated control of tubulin synthesis in animal cells. Curr Opin Cell Biol. 1989;1(1):10-14. doi: 10.1016/S0955-0674(89)80031-9
Steinmetz MO, Prota AE. Microtubule-Targeting Agents: Strategies To Hijack the Cytoskeleton. Trends Cell Biol. 2018;28:776-792. doi: 10.1016/j.tcb.2018.05.001
Steinmetz MO, Prota AE. Structure-based discovery and rational design of microtubule-targeting agents. Curr Opin Struct Biol. 2024;87:102845. doi: 10.1016/j.sbi.2024.102845
Ben-Ze’ev A, Farmer SR, Penman S. Mechanisms of regulating tubulin synthesis in cultured mammalian cells. Cell. 1979;17:319-325. doi: 10.1016/0092-8674(79)90193-1
Yen TJ, Machlin PS, Cleveland DW. Autoregulated instability of beta-tubulin mRNAs by recognition of the nascent amino terminus of beta-tubulin. Nature. 1988;334:580-585. doi: 10.1038/334580a0
Lin Z, et al. TTC5 mediates autoregulation of tubulin via mRNA degradation. Science. 2020;367:100-104. doi: 10.1126/science.aaz5891
Hopfler M, et al. Mechanism of ribosome-associated mRNA degradation during tubulin autoregulation. Mol Cell. 2023;83:2290-2302. doi: 10.1016/j.molcel.2023.06.027
Batiuk A, et al. Soluble alphabeta-tubulins reversibly sequester TTC5 to regulate tubulin mRNA decay. Nat Commun. 2024;15:9963. doi: 10.1038/s41467-024-54205-1
Yu N, et al. Isolation of Functional Tubulin Dimers and of Tubulin-Associated Proteins from Mammalian Cells. Curr Biol. 2016;26:1728-1736. doi: 10.1016/j.cub.2016.05.018
Yu N, Galjart N. TAPping into the treasures of tubulin using novel protein production methods. Essays Biochem. 2018;62:781-792. doi: 10.1042/ebc20180036
Urlinger S, et al. Exploring the sequence space for tetracycline-dependent transcriptional activators: novel mutations yield expanded range and sensitivity. Proc Natl Acad Sci U S A. 2000;97:7963-7968. doi: 10.1073/pnas.130192697
Aillaud C, et al. Vasohibins/SVBP are tubulin carboxypeptidases (TCPs) that regulate neuron differentiation. Science. 2017;358:1448-1453. doi: 10.1126/science.aao4147
L’Hernault SW, Rosenbaum JL. Chlamydomonas alpha-tubulin is posttranslationally modified in the flagella during flagellar assembly. J Cell Biol. 1983;97:258-263. doi: 10.1083/jcb.97.1.258
Nieuwenhuis J, et al. Vasohibins encode tubulin detyrosinating activity. Science. 2017;358:1453-1456. doi: 10.1126/science.aao5891
Schroder HC, Wehland J, Weber K. Purification of brain tubulin-tyrosine ligase by biochemical and immunological methods. J Cell Biol. 1985;100:276-281. doi: 10.1083/jcb.100.1.276
Hammond JW, et al. Posttranslational modifications of tubulin and the polarized transport of kinesin-1 in neurons. Mol Biol Cell. 2010;21:572-583. doi: 10.1091/mbc.e09-11-0909
Stepanova T, et al. Visualization of microtubule growth in cultured neurons via the use of EB3-GFP (end-binding protein 3-green fluorescent protein). J Neurosci. 2003;23:2655-2664. doi: 10.1523/JNEUROSCI.23-07-02655.2003
Wood LM, Moore JK. beta3 accelerates microtubule plus end maturation through a divergent lateral interface. Mol Biol Cell. 2025;36:ar36. doi: 10.1091/mbc.e24-09-0450
Kok M, Huber F, Kalisch SM, Dogterom M. EB3-informed dynamics of the microtubule stabilizing cap during stalled growth. Biophys J. 2024. doi: 10.1016/j.bpj.2024.03.016
Cho NH, et al. OpenCell: Endogenous tagging for the cartography of human cellular organization. Science. 2022;375:eabi6983. doi: 10.1126/science.abi6983
Folker ES, Baker BM, Goodson HV. Interactions between CLIP-170, tubulin, and microtubules: implications for the mechanism of Clip-170 plus-end tracking behavior. Mol Biol Cell. 2005;16:5373-5384. doi: 10.1091/mbc.e05-05-0461
Honnappa S, et al. Key interaction modes of dynamic +TIP networks. Mol Cell. 2006;23:663-671. doi: 10.1016/j.molcel.2006.07.013
Mishima M, et al. Structural basis for tubulin recognition by cytoplasmic linker protein 170 and its autoinhibition. Proc Natl Acad Sci U S A. 2007;104:10346-10351. doi: 10.1073/pnas.0703189104
Boyko S, Li Q, Surewicz K, Surewicz WK. Distinct liquid-liquid phase separation properties of end-binding proteins EB1 and EB3. J Biol Chem. 2025;110849. doi: 10.1016/j.jbc.2024.110849
Meier SM, et al. Multivalency ensures persistence of a +TIP body at specialized microtubule ends. Nat Cell Biol. 2023;25:56-67. doi: 10.1038/s41556-022-01010-z
Meier SM, Steinmetz MO, Barral Y. Microtubule specialization by +TIP networks: from mechanisms to functional implications. Trends Biochem Sci. 2024;49:318-332. doi: 10.1016/j.tibs.2024.02.001
Borisy G, et al. Microtubules: 50 years on from the discovery of tubulin. Nat Rev Mol Cell Biol. 2016;17:322-328. doi: 10.1038/nrm.2016.37
Hosea R, Hillary S, Naqvi S, Wu S, Kasim V. The two sides of chromosomal instability: drivers and brakes in cancer. Signal Transduct Target Ther. 2024;9:75. doi: 10.1038/s41392-024-01795-1
Schmidt M, Medema RH. Exploiting the compromised spindle assembly checkpoint function of tumor cells: dawn on the horizon? Cell Cycle. 2006;5:159-163. doi: 10.4161/cc.5.2.2385
Lukinavicius G, et al. Fluorogenic probes for live-cell imaging of the cytoskeleton. Nat Methods. 2014;11:731-733. doi: 10.1038/nmeth.2972
Tomer R, Khairy K, Keller PJ. Light sheet microscopy in cell biology. Methods Mol Biol. 2013;931:123-137. doi: 10.1007/978-1-62703-056-4_6
Musacchio A, Salmon ED. The spindle-assembly checkpoint in space and time. Nat Rev Mol Cell Biol. 2007;8:379-393. doi: 10.1038/nrm2163
Jordan MA, Thrower D, Wilson L. Effects of vinblastine, podophyllotoxin and nocodazole on mitotic spindles. Implications for the role of microtubule dynamics in mitosis. J Cell Sci. 1992;102(Pt 3):401-416. doi: 10.1242/jcs.102.3.401
Jordan MA, Toso RJ, Thrower D, Wilson L. Mechanism of mitotic block and inhibition of cell proliferation by taxol at low concentrations. Proc Natl Acad Sci U S A. 1993;90:9552-9556. doi: 10.1073/pnas.90.20.9552
Sudakin V, Chan GK, Yen TJ. Checkpoint inhibition of the APC/C in HeLa cells is mediated by a complex of BUBR1, BUB3, CDC20, and MAD2. J Cell Biol. 2001;154:925-936. doi: 10.1083/jcb.154.5.925
Logarinho E, Bousbaa H. Kinetochore-microtubule interactions “in check” by Bub1, Bub3 and BubR1: The dual task of attaching and signalling. Cell Cycle. 2008;7:1763-1768. doi: 10.4161/cc.7.11.6006
Gaetz J, Kapoor TM. Dynein/dynactin regulate metaphase spindle length by targeting depolymerizing activities to spindle poles. J Cell Biol. 2004;166:465-471. doi: 10.1083/jcb.200404096
Ganem NJ, Compton DA. The KinI kinesin Kif2a is required for bipolar spindle assembly through a functional relationship with MCAK. J Cell Biol. 2004;166:473-478. doi: 10.1083/jcb.200404097
Uehara R, et al. Aurora B and Kif2A control microtubule length for assembly of a functional central spindle during anaphase. J Cell Biol. 2013;202:623-636. doi: 10.1083/jcb.201302123
Girao H, et al. CLASP2 binding to curved microtubule tips promotes flux and stabilizes kinetochore attachments. J Cell Biol. 2020;219(12):e202005137. doi: 10.1083/jcb.202005137
Logarinho E, et al. CLASPs prevent irreversible multipolarity by ensuring spindle-pole resistance to traction forces during chromosome alignment. Nat Cell Biol. 2012;14:295-303. doi: 10.1038/ncb2427
Maffini S, et al. Motor-independent targeting of CLASPs to kinetochores by CENP-E promotes microtubule turnover and poleward flux. Curr Biol. 2009;19:1566-1572. doi: 10.1016/j.cub.2009.07.056
Zhai Y, Kronebusch PJ, Simon PM, Borisy GG. Microtubule dynamics at the G2/M transition: abrupt breakdown of cytoplasmic microtubules at nuclear envelope breakdown and implications for spindle morphogenesis. J Cell Biol. 1996;135:201-214. doi: 10.1083/jcb.135.1.201
Zhai Y, Borisy GG. Quantitative determination of the proportion of microtubule polymer present during the mitosis-interphase transition. J Cell Sci. 1994;107(Pt 4):881-890. doi: 10.1242/jcs.107.4.881
Travis SM, Mahon BP, Petry S. How Microtubules Build the Spindle Branch by Branch. Annu Rev Cell Dev Biol. 2022;38:1-23. doi: 10.1146/annurev-cellbio-120221-095129
Zhou AS, et al. Diverse microtubule-targeted anticancer agents kill cells by inducing chromosome missegregation on multipolar spindles. PLoS Biol. 2023;21:e3002339. doi: 10.1371/journal.pbio.3002339
Helin K. Regulation of cell proliferation by the E2F transcription factors. Curr Opin Genet Dev. 1998;8:28-35. doi: 10.1016/S0959-437X(98)80006-2
Henley SA, Dick FA. The retinoblastoma family of proteins and their regulatory functions in the mammalian cell division cycle. Cell Div. 2012;7:10. doi: 10.1186/1747-1028-7-10
Weinberg RA. The retinoblastoma protein and cell cycle control. Cell. 1995;81:323-330. doi: 10.1016/0092-8674(95)90385-2
Rogakou EP, Pilch DR, Orr AH, Ivanova VS, Bonner WM. DNA double-stranded breaks induce histone H2AX phosphorylation on serine 139. J Biol Chem. 1998;273:5858-5868. doi: 10.1074/jbc.273.10.5858
Sedelnikova OA, Pilch DR, Redon C, Bonner WM. Histone H2AX in DNA damage and repair. Cancer Biol Ther. 2003;2:233-235. doi: 10.4161/cbt.2.3.633
Mazumdar M, Sung MH, Misteli T. Chromatin maintenance by a molecular motor protein. Nucleus. 2011;2:591-600. doi: 10.4161/nucl.2.6.18214
Dullovi A, et al. Microtubule-associated proteins MAP7 and MAP7D1 promote DNA double-strand break repair in the G1 cell cycle phase. iScience. 2023;26:106107. doi: 10.1016/j.isci.2023.106107
Luessing J, et al. The nuclear kinesin KIF18B promotes 53BP1-mediated DNA double-strand break repair. Cell Rep. 2021;35:109306. doi: 10.1016/j.celrep.2021.109306
Zhu S, et al. Kinesin Kif2C in regulation of DNA double strand break dynamics and repair. Elife. 2020;9:e56520. doi: 10.7554/eLife.56520
Ma S, et al. DNA damage promotes microtubule dynamics through a DNA-PK-AKT axis for enhanced repair. J Cell Biol. 2021;220(4):e202008094. doi: 10.1083/jcb.202008094
Liu P, Wurtz M, Zupa E, Pfeffer S, Schiebel E. Microtubule nucleation: The waltz between gamma-tubulin ring complex and associated proteins. Curr Opin Cell Biol. 2021;68:124-131. doi: 10.1016/j.ceb.2020.10.002
Gasic I, Boswell SA, Mitchison TJ. Tubulin mRNA stability is sensitive to change in microtubule dynamics caused by multiple physiological and toxic cues. PLoS Biol. 2019;17:e3000225. doi: 10.1371/journal.pbio.3000225
Assaf M, Lacheheub C, Gasic I. Tubulin autoregulation controls the biosynthesis of γ-tubulin to ensure mitotic fidelity. bioRxiv. 2026. doi: 10.1101/2026.01.01.716648
Zhou Y, et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun. 2019;10:1523. doi: 10.1038/s41467-019-09234-6
Liberzon A, et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 2015;1:417-425. doi: 10.1016/j.cels.2015.12.004
Subramanian A, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005;102:15545-15550. doi: 10.1073/pnas.0506580102
Jordan MA, Kamath K. How do microtubule-targeted drugs work? An overview. Curr Cancer Drug Targets. 2007;7:730-742. doi: 10.2174/156800907783220534
Semenza GL, Shimoda LA, Prabhakar NR. Regulation of gene expression by HIF1. Novartis Found Symp. 2006;272:2-8. doi: 10.1002/0470023678.ch1
Yfantis A, et al. Transcriptional Response to Hypoxia: The Role of HIF-1-Associated Co-Regulators. Cells. 2023;12(3):471. doi: 10.3390/cells12030471
Costa-Mattioli M, Walter P. The integrated stress response: From mechanism to disease. Science. 2020;368(6489):eaat5314. doi: 10.1126/science.aat5314
Chun Y, Kim J. AMPK-mTOR Signaling and Cellular Adaptations in Hypoxia. Int J Mol Sci. 2021;22(8):4077. doi: 10.3390/ijms22084077
Harding HP, et al. An integrated stress response regulates amino acid metabolism and resistance to oxidative stress